GPy.core package¶
Introduction¶
This module contains the fundamental classes of GPy - classes that are inherited by objects in other parts of GPy in order to provide a consistent interface to major functionality.

GPy.core.model is inherited by
GPy.core.gp.GP. And GPy.core.model itself
inherits paramz.model.Model from the paramz
package. paramz essentially provides an inherited set of properties
and functions used to manage state (and state changes) of the model.
GPy.core.gp.GP represents a GP model. Such an entity is
typically passed variables representing known (x) and observed (y)
data, along with a kernel and other information needed to create the
specific model. It exposes functions which return information derived
from the inputs to the model, for example predicting unobserved
variables based on new known variables, or the log marginal likelihood
of the current state of the model.
optimize is called to optimize
hyperparameters of the model. The optimizer argument takes a string
which is used to specify non-default optimization schemes.
Various plotting functions can be called against GPy.core.gp.GP.
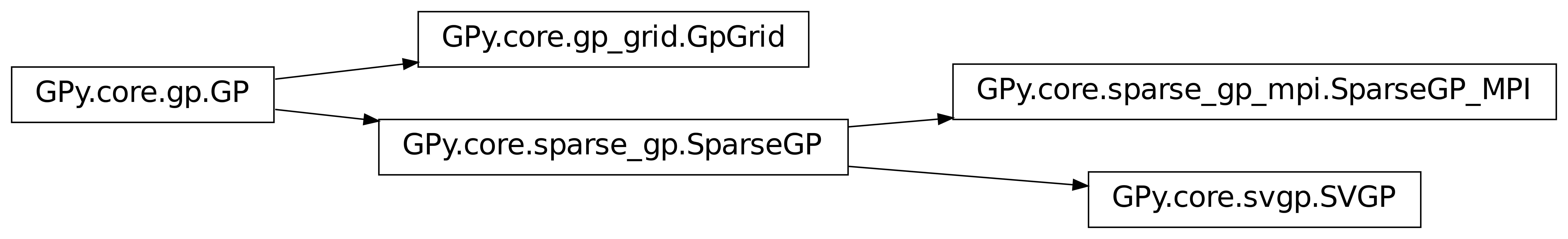
GPy.core.gp.GP is used as the basis for classes supporting
more specialized types of Gaussian Process model. These are however
generally still not specific enough to be called by the user and are
inhereted by members of the GPy.models package.
-
randomize(self, rand_gen=None, *args, **kwargs)[source]¶ Randomize the model. Make this draw from the prior if one exists, else draw from given random generator
Parameters: - rand_gen – np random number generator which takes args and kwargs
- loc (flaot) – loc parameter for random number generator
- scale (float) – scale parameter for random number generator
- kwargs (args,) – will be passed through to random number generator
Subpackages¶
Submodules¶
GPy.core.gp module¶
-
class
GP(X, Y, kernel, likelihood, mean_function=None, inference_method=None, name='gp', Y_metadata=None, normalizer=False)[source]¶ Bases:
GPy.core.model.ModelGeneral purpose Gaussian process model
Parameters: - X – input observations
- Y – output observations
- kernel – a GPy kernel
- likelihood – a GPy likelihood
- inference_method – The
LatentFunctionInferenceinference method to use for this GP - normalizer (Norm) – normalize the outputs Y. Prediction will be un-normalized using this normalizer. If normalizer is True, we will normalize using Standardize. If normalizer is False, no normalization will be done.
Return type: model object
Note
Multiple independent outputs are allowed using columns of Y
-
infer_newX(Y_new, optimize=True)[source]¶ Infer X for the new observed data Y_new.
Parameters: - Y_new (numpy.ndarray) – the new observed data for inference
- optimize (boolean) – whether to optimize the location of new X (True by default)
Returns: a tuple containing the posterior estimation of X and the model that optimize X
Return type: (
VariationalPosteriorand numpy.ndarray,Model)
-
log_likelihood()[source]¶ The log marginal likelihood of the model, \(p(\mathbf{y})\), this is the objective function of the model being optimised
-
log_predictive_density(x_test, y_test, Y_metadata=None)[source]¶ Calculation of the log predictive density
Parameters: - x_test ((Nx1) array) – test locations (x_{*})
- y_test ((Nx1) array) – test observations (y_{*})
- Y_metadata – metadata associated with the test points
-
log_predictive_density_sampling(x_test, y_test, Y_metadata=None, num_samples=1000)[source]¶ Calculation of the log predictive density by sampling
Parameters: - x_test ((Nx1) array) – test locations (x_{*})
- y_test ((Nx1) array) – test observations (y_{*})
- Y_metadata – metadata associated with the test points
- num_samples (int) – number of samples to use in monte carlo integration
-
optimize(optimizer=None, start=None, messages=False, max_iters=1000, ipython_notebook=True, clear_after_finish=False, **kwargs)[source]¶ Optimize the model using self.log_likelihood and self.log_likelihood_gradient, as well as self.priors. kwargs are passed to the optimizer. They can be:
Parameters: - max_iters (int) – maximum number of function evaluations
- messages (bool) – whether to display during optimisation
- optimizer (string) – which optimizer to use (defaults to self.preferred optimizer), a range of optimisers can be found in :module:`~GPy.inference.optimization`, they include ‘scg’, ‘lbfgs’, ‘tnc’.
- ipython_notebook (bool) – whether to use ipython notebook widgets or not.
- clear_after_finish (bool) – if in ipython notebook, we can clear the widgets after optimization.
-
parameters_changed()[source]¶ Method that is called upon any changes to
Paramvariables within the model. In particular in the GP class this method re-performs inference, recalculating the posterior and log marginal likelihood and gradients of the modelWarning
This method is not designed to be called manually, the framework is set up to automatically call this method upon changes to parameters, if you call this method yourself, there may be unexpected consequences.
-
plot(plot_limits=None, fixed_inputs=None, resolution=None, plot_raw=False, apply_link=False, which_data_ycols='all', which_data_rows='all', visible_dims=None, levels=20, samples=0, samples_likelihood=0, lower=2.5, upper=97.5, plot_data=True, plot_inducing=True, plot_density=False, predict_kw=None, projection='2d', legend=True, **kwargs)¶ Convenience function for plotting the fit of a GP.
You can deactivate the legend for this one plot by supplying None to label.
Give the Y_metadata in the predict_kw if you need it.
If you want fine graned control use the specific plotting functions supplied in the model.
Parameters: - plot_limits (np.array) – The limits of the plot. If 1D [xmin,xmax], if 2D [[xmin,ymin],[xmax,ymax]]. Defaluts to data limits
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- resolution (int) – The resolution of the prediction [default:200]
- plot_raw (bool) – plot the latent function (usually denoted f) only?
- apply_link (bool) – whether to apply the link function of the GP to the raw prediction.
- which_data_ycols ('all' or a list of integers) – when the data has several columns (independant outputs), only plot these
- which_data_rows ('all' or a slice object to slice self.X, self.Y) – which of the training data to plot (default all)
- visible_dims (array-like) – which columns of the input X (!) to plot (array-like or list of ints)
- levels (int) – the number of levels in the density (number bigger then 1, where 35 is smooth and 1 is the same as plot_confidence). You can go higher then 50 if the result is not smooth enough for you.
- samples (int) – the number of samples to draw from the GP and plot into the plot. This will allways be samples from the latent function.
- samples_likelihood (int) – the number of samples to draw from the GP and apply the likelihood noise. This is usually not what you want!
- lower (float) – the lower percentile to plot
- upper (float) – the upper percentile to plot
- plot_data (bool) – plot the data into the plot?
- plot_inducing (bool) – plot inducing inputs?
- plot_density (bool) – plot density instead of the confidence interval?
- predict_kw (dict) – the keyword arguments for the prediction. If you want to plot a specific kernel give dict(kern=<specific kernel>) in here
- projection ({2d|3d}) – plot in 2d or 3d?
- legend (bool) – convenience, whether to put a legend on the plot or not.
-
plot_confidence(lower=2.5, upper=97.5, plot_limits=None, fixed_inputs=None, resolution=None, plot_raw=False, apply_link=False, visible_dims=None, which_data_ycols='all', label='gp confidence', predict_kw=None, **kwargs)¶ Plot the confidence interval between the percentiles lower and upper. E.g. the 95% confidence interval is $2.5, 97.5$. Note: Only implemented for one dimension!
You can deactivate the legend for this one plot by supplying None to label.
Give the Y_metadata in the predict_kw if you need it.
Parameters: - lower (float) – the lower percentile to plot
- upper (float) – the upper percentile to plot
- plot_limits (np.array) – The limits of the plot. If 1D [xmin,xmax], if 2D [[xmin,ymin],[xmax,ymax]]. Defaluts to data limits
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- resolution (int) – The resolution of the prediction [default:200]
- plot_raw (bool) – plot the latent function (usually denoted f) only?
- apply_link (bool) – whether to apply the link function of the GP to the raw prediction.
- visible_dims (array-like) – which columns of the input X (!) to plot (array-like or list of ints)
- which_data_ycols (array-like) – which columns of the output y (!) to plot (array-like or list of ints)
- predict_kw (dict) – the keyword arguments for the prediction. If you want to plot a specific kernel give dict(kern=<specific kernel>) in here
-
plot_data(which_data_rows='all', which_data_ycols='all', visible_dims=None, projection='2d', label=None, **plot_kwargs)¶ - Plot the training data
- For higher dimensions than two, use fixed_inputs to plot the data points with some of the inputs fixed.
Can plot only part of the data using which_data_rows and which_data_ycols.
Parameters: - which_data_rows ('all' or a slice object to slice self.X, self.Y) – which of the training data to plot (default all)
- which_data_ycols ('all' or a list of integers) – when the data has several columns (independant outputs), only plot these
- visible_dims (a numpy array) – an array specifying the input dimensions to plot (maximum two)
- projection ({'2d','3d'}) – whether to plot in 2d or 3d. This only applies when plotting two dimensional inputs!
- label (str) – the label for the plot
- plot_kwargs (kwargs) – kwargs for the data plot for the plotting library you are using
Returns list: of plots created.
-
plot_data_error(which_data_rows='all', which_data_ycols='all', visible_dims=None, projection='2d', label=None, **error_kwargs)¶ Plot the training data input error.
For higher dimensions than two, use fixed_inputs to plot the data points with some of the inputs fixed.
Can plot only part of the data using which_data_rows and which_data_ycols.
Parameters: - which_data_rows ('all' or a slice object to slice self.X, self.Y) – which of the training data to plot (default all)
- which_data_ycols ('all' or a list of integers) – when the data has several columns (independant outputs), only plot these
- visible_dims (a numpy array) – an array specifying the input dimensions to plot (maximum two)
- projection ({'2d','3d'}) – whether to plot in 2d or 3d. This only applies when plotting two dimensional inputs!
- error_kwargs (dict) – kwargs for the error plot for the plotting library you are using
- label (str) – the label for the plot
- plot_kwargs (kwargs) – kwargs for the data plot for the plotting library you are using
Returns list: of plots created.
-
plot_density(plot_limits=None, fixed_inputs=None, resolution=None, plot_raw=False, apply_link=False, visible_dims=None, which_data_ycols='all', levels=35, label='gp density', predict_kw=None, **kwargs)¶ Plot the confidence interval between the percentiles lower and upper. E.g. the 95% confidence interval is $2.5, 97.5$. Note: Only implemented for one dimension!
You can deactivate the legend for this one plot by supplying None to label.
Give the Y_metadata in the predict_kw if you need it.
Parameters: - plot_limits (np.array) – The limits of the plot. If 1D [xmin,xmax], if 2D [[xmin,ymin],[xmax,ymax]]. Defaluts to data limits
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- resolution (int) – The resolution of the prediction [default:200]
- plot_raw (bool) – plot the latent function (usually denoted f) only?
- apply_link (bool) – whether to apply the link function of the GP to the raw prediction.
- visible_dims (array-like) – which columns of the input X (!) to plot (array-like or list of ints)
- which_data_ycols (array-like) – which columns of y to plot (array-like or list of ints)
- levels (int) – the number of levels in the density (number bigger then 1, where 35 is smooth and 1 is the same as plot_confidence). You can go higher then 50 if the result is not smooth enough for you.
- predict_kw (dict) – the keyword arguments for the prediction. If you want to plot a specific kernel give dict(kern=<specific kernel>) in here
-
plot_errorbars_trainset(which_data_rows='all', which_data_ycols='all', fixed_inputs=None, plot_raw=False, apply_link=False, label=None, projection='2d', predict_kw=None, **plot_kwargs)¶ Plot the errorbars of the GP likelihood on the training data. These are the errorbars after the appropriate approximations according to the likelihood are done.
This also works for heteroscedastic likelihoods.
Give the Y_metadata in the predict_kw if you need it.
Parameters: - which_data_rows ('all' or a slice object to slice self.X, self.Y) – which of the training data to plot (default all)
- which_data_ycols – when the data has several columns (independant outputs), only plot these
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- predict_kwargs (dict) – kwargs for the prediction used to predict the right quantiles.
- plot_kwargs (kwargs) – kwargs for the data plot for the plotting library you are using
-
plot_f(plot_limits=None, fixed_inputs=None, resolution=None, apply_link=False, which_data_ycols='all', which_data_rows='all', visible_dims=None, levels=20, samples=0, lower=2.5, upper=97.5, plot_density=False, plot_data=True, plot_inducing=True, projection='2d', legend=True, predict_kw=None, **kwargs)¶ Convinience function for plotting the fit of a GP. This is the same as plot, except it plots the latent function fit of the GP!
If you want fine graned control use the specific plotting functions supplied in the model.
You can deactivate the legend for this one plot by supplying None to label.
Give the Y_metadata in the predict_kw if you need it.
Parameters: - plot_limits (np.array) – The limits of the plot. If 1D [xmin,xmax], if 2D [[xmin,ymin],[xmax,ymax]]. Defaluts to data limits
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- resolution (int) – The resolution of the prediction [default:200]
- apply_link (bool) – whether to apply the link function of the GP to the raw prediction.
- which_data_ycols ('all' or a list of integers) – when the data has several columns (independant outputs), only plot these
- which_data_rows ('all' or a slice object to slice self.X, self.Y) – which of the training data to plot (default all)
- visible_dims (array-like) – an array specifying the input dimensions to plot (maximum two)
- levels (int) – the number of levels in the density (number bigger then 1, where 35 is smooth and 1 is the same as plot_confidence). You can go higher then 50 if the result is not smooth enough for you.
- samples (int) – the number of samples to draw from the GP and plot into the plot. This will allways be samples from the latent function.
- lower (float) – the lower percentile to plot
- upper (float) – the upper percentile to plot
- plot_data (bool) – plot the data into the plot?
- plot_inducing (bool) – plot inducing inputs?
- plot_density (bool) – plot density instead of the confidence interval?
- predict_kw (dict) – the keyword arguments for the prediction. If you want to plot a specific kernel give dict(kern=<specific kernel>) in here
- error_kwargs (dict) – kwargs for the error plot for the plotting library you are using
- plot_kwargs (kwargs) – kwargs for the data plot for the plotting library you are using
-
plot_latent(plot_limits=None, fixed_inputs=None, resolution=None, apply_link=False, which_data_ycols='all', which_data_rows='all', visible_dims=None, levels=20, samples=0, lower=2.5, upper=97.5, plot_density=False, plot_data=True, plot_inducing=True, projection='2d', legend=True, predict_kw=None, **kwargs)¶ Convinience function for plotting the fit of a GP. This is the same as plot, except it plots the latent function fit of the GP!
If you want fine graned control use the specific plotting functions supplied in the model.
You can deactivate the legend for this one plot by supplying None to label.
Give the Y_metadata in the predict_kw if you need it.
Parameters: - plot_limits (np.array) – The limits of the plot. If 1D [xmin,xmax], if 2D [[xmin,ymin],[xmax,ymax]]. Defaluts to data limits
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- resolution (int) – The resolution of the prediction [default:200]
- apply_link (bool) – whether to apply the link function of the GP to the raw prediction.
- which_data_ycols ('all' or a list of integers) – when the data has several columns (independant outputs), only plot these
- which_data_rows ('all' or a slice object to slice self.X, self.Y) – which of the training data to plot (default all)
- visible_dims (array-like) – an array specifying the input dimensions to plot (maximum two)
- levels (int) – the number of levels in the density (number bigger then 1, where 35 is smooth and 1 is the same as plot_confidence). You can go higher then 50 if the result is not smooth enough for you.
- samples (int) – the number of samples to draw from the GP and plot into the plot. This will allways be samples from the latent function.
- lower (float) – the lower percentile to plot
- upper (float) – the upper percentile to plot
- plot_data (bool) – plot the data into the plot?
- plot_inducing (bool) – plot inducing inputs?
- plot_density (bool) – plot density instead of the confidence interval?
- predict_kw (dict) – the keyword arguments for the prediction. If you want to plot a specific kernel give dict(kern=<specific kernel>) in here
- error_kwargs (dict) – kwargs for the error plot for the plotting library you are using
- plot_kwargs (kwargs) – kwargs for the data plot for the plotting library you are using
-
plot_magnification(labels=None, which_indices=None, resolution=60, marker='<>^vsd', legend=True, plot_limits=None, updates=False, mean=True, covariance=True, kern=None, num_samples=1000, scatter_kwargs=None, plot_scatter=True, **imshow_kwargs)¶ Plot the magnification factor of the GP on the inputs. This is the density of the GP as a gray scale.
Parameters: - labels (array-like) – a label for each data point (row) of the inputs
- which_indices ((int, int)) – which input dimensions to plot against each other
- resolution (int) – the resolution at which we predict the magnification factor
- marker (str) – markers to use - cycle if more labels then markers are given
- legend (bool) – whether to plot the legend on the figure
- plot_limits ((xmin, xmax, ymin, ymax) or ((xmin, xmax), (ymin, ymax))) – the plot limits for the plot
- updates (bool) – if possible, make interactive updates using the specific library you are using
- mean (bool) – use the mean of the Wishart embedding for the magnification factor
- covariance (bool) – use the covariance of the Wishart embedding for the magnification factor
- kern (
Kern) – the kernel to use for prediction - num_samples (int) – the number of samples to plot maximally. We do a stratified subsample from the labels, if the number of samples (in X) is higher then num_samples.
- imshow_kwargs – the kwargs for the imshow (magnification factor)
- kwargs – the kwargs for the scatter plots
-
plot_mean(plot_limits=None, fixed_inputs=None, resolution=None, plot_raw=False, apply_link=False, visible_dims=None, which_data_ycols='all', levels=20, projection='2d', label='gp mean', predict_kw=None, **kwargs)¶ Plot the mean of the GP.
You can deactivate the legend for this one plot by supplying None to label.
Give the Y_metadata in the predict_kw if you need it.
Parameters: - plot_limits (np.array) – The limits of the plot. If 1D [xmin,xmax], if 2D [[xmin,ymin],[xmax,ymax]]. Defaluts to data limits
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- resolution (int) – The resolution of the prediction [defaults are 1D:200, 2D:50]
- plot_raw (bool) – plot the latent function (usually denoted f) only?
- apply_link (bool) – whether to apply the link function of the GP to the raw prediction.
- which_data_ycols (array-like) – which columns of y to plot (array-like or list of ints)
- levels (int) – for 2D plotting, the number of contour levels to use is
- projection ({'2d','3d'}) – whether to plot in 2d or 3d. This only applies when plotting two dimensional inputs!
- label (str) – the label for the plot.
- predict_kw (dict) – the keyword arguments for the prediction. If you want to plot a specific kernel give dict(kern=<specific kernel>) in here
-
plot_noiseless(plot_limits=None, fixed_inputs=None, resolution=None, apply_link=False, which_data_ycols='all', which_data_rows='all', visible_dims=None, levels=20, samples=0, lower=2.5, upper=97.5, plot_density=False, plot_data=True, plot_inducing=True, projection='2d', legend=True, predict_kw=None, **kwargs)¶ Convinience function for plotting the fit of a GP. This is the same as plot, except it plots the latent function fit of the GP!
If you want fine graned control use the specific plotting functions supplied in the model.
You can deactivate the legend for this one plot by supplying None to label.
Give the Y_metadata in the predict_kw if you need it.
Parameters: - plot_limits (np.array) – The limits of the plot. If 1D [xmin,xmax], if 2D [[xmin,ymin],[xmax,ymax]]. Defaluts to data limits
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- resolution (int) – The resolution of the prediction [default:200]
- apply_link (bool) – whether to apply the link function of the GP to the raw prediction.
- which_data_ycols ('all' or a list of integers) – when the data has several columns (independant outputs), only plot these
- which_data_rows ('all' or a slice object to slice self.X, self.Y) – which of the training data to plot (default all)
- visible_dims (array-like) – an array specifying the input dimensions to plot (maximum two)
- levels (int) – the number of levels in the density (number bigger then 1, where 35 is smooth and 1 is the same as plot_confidence). You can go higher then 50 if the result is not smooth enough for you.
- samples (int) – the number of samples to draw from the GP and plot into the plot. This will allways be samples from the latent function.
- lower (float) – the lower percentile to plot
- upper (float) – the upper percentile to plot
- plot_data (bool) – plot the data into the plot?
- plot_inducing (bool) – plot inducing inputs?
- plot_density (bool) – plot density instead of the confidence interval?
- predict_kw (dict) – the keyword arguments for the prediction. If you want to plot a specific kernel give dict(kern=<specific kernel>) in here
- error_kwargs (dict) – kwargs for the error plot for the plotting library you are using
- plot_kwargs (kwargs) – kwargs for the data plot for the plotting library you are using
-
plot_samples(plot_limits=None, fixed_inputs=None, resolution=None, plot_raw=True, apply_link=False, visible_dims=None, which_data_ycols='all', samples=3, projection='2d', label='gp_samples', predict_kw=None, **kwargs)¶ Plot the mean of the GP.
You can deactivate the legend for this one plot by supplying None to label.
Give the Y_metadata in the predict_kw if you need it.
Parameters: - plot_limits (np.array) – The limits of the plot. If 1D [xmin,xmax], if 2D [[xmin,ymin],[xmax,ymax]]. Defaluts to data limits
- fixed_inputs (a list of tuples) – a list of tuple [(i,v), (i,v)…], specifying that input dimension i should be set to value v.
- resolution (int) – The resolution of the prediction [defaults are 1D:200, 2D:50]
- plot_raw (bool) – plot the latent function (usually denoted f) only? This is usually what you want!
- apply_link (bool) – whether to apply the link function of the GP to the raw prediction.
- visible_dims (array-like) – which columns of the input X (!) to plot (array-like or list of ints)
- which_data_ycols (array-like) – which columns of y to plot (array-like or list of ints)
- predict_kw (dict) – the keyword arguments for the prediction. If you want to plot a specific kernel give dict(kern=<specific kernel>) in here
- levels (int) – for 2D plotting, the number of contour levels to use is
-
posterior_covariance_between_points(X1, X2, Y_metadata=None, likelihood=None, include_likelihood=True)[source]¶ Computes the posterior covariance between points. Includes likelihood variance as well as normalization so that evaluation at (x,x) is consistent with model.predict
Parameters: - X1 – some input observations
- X2 – other input observations
- Y_metadata – metadata about the predicting point to pass to the likelihood
- include_likelihood (bool) – Whether or not to add likelihood noise to the predicted underlying latent function f.
Returns: cov: posterior covariance, a Numpy array, Nnew x Nnew if self.output_dim == 1, and Nnew x Nnew x self.output_dim otherwise.
-
posterior_samples(X, size=10, Y_metadata=None, likelihood=None, **predict_kwargs)[source]¶ Samples the posterior GP at the points X.
Parameters: - X (np.ndarray (Nnew x self.input_dim.)) – the points at which to take the samples.
- size (int.) – the number of a posteriori samples.
- noise_model (integer.) – for mixed noise likelihood, the noise model to use in the samples.
Returns: Ysim: set of simulations,
Return type: np.ndarray (D x N x samples) (if D==1 we flatten out the first dimension)
-
posterior_samples_f(X, size=10, **predict_kwargs)[source]¶ Samples the posterior GP at the points X.
Parameters: - X (np.ndarray (Nnew x self.input_dim)) – The points at which to take the samples.
- size (int.) – the number of a posteriori samples.
Returns: set of simulations
Return type: np.ndarray (Nnew x D x samples)
-
predict(Xnew, full_cov=False, Y_metadata=None, kern=None, likelihood=None, include_likelihood=True)[source]¶ Predict the function(s) at the new point(s) Xnew. This includes the likelihood variance added to the predicted underlying function (usually referred to as f).
In order to predict without adding in the likelihood give include_likelihood=False, or refer to self.predict_noiseless().
Parameters: - Xnew (np.ndarray (Nnew x self.input_dim)) – The points at which to make a prediction
- full_cov (bool) – whether to return the full covariance matrix, or just the diagonal
- Y_metadata – metadata about the predicting point to pass to the likelihood
- kern – The kernel to use for prediction (defaults to the model kern). this is useful for examining e.g. subprocesses.
- include_likelihood (bool) – Whether or not to add likelihood noise to the predicted underlying latent function f.
Returns: (mean, var): mean: posterior mean, a Numpy array, Nnew x self.input_dim var: posterior variance, a Numpy array, Nnew x 1 if full_cov=False,
Nnew x Nnew otherwise
If full_cov and self.input_dim > 1, the return shape of var is Nnew x Nnew x self.input_dim. If self.input_dim == 1, the return shape is Nnew x Nnew. This is to allow for different normalizations of the output dimensions.
Note: If you want the predictive quantiles (e.g. 95% confidence interval) use
predict_quantiles.
-
predict_jacobian(Xnew, kern=None, full_cov=False)[source]¶ Compute the derivatives of the posterior of the GP.
Given a set of points at which to predict X* (size [N*,Q]), compute the mean and variance of the derivative. Resulting arrays are sized:
- dL_dX* – [N*, Q ,D], where D is the number of output in this GP (usually one).
- Note that this is the mean and variance of the derivative, not the derivative of the mean and variance! (See predictive_gradients for that)
- dv_dX* – [N*, Q], (since all outputs have the same variance)
- If there is missing data, it is not implemented for now, but there will be one output variance per output dimension.
Parameters: - X (np.ndarray (Xnew x self.input_dim)) – The points at which to get the predictive gradients.
- kern – The kernel to compute the jacobian for.
- full_cov (boolean) – whether to return the cross-covariance terms between
the N* Jacobian vectors
Returns: dmu_dX, dv_dX Return type: [np.ndarray (N*, Q ,D), np.ndarray (N*,Q,(D)) ]
-
predict_magnification(Xnew, kern=None, mean=True, covariance=True, dimensions=None)[source]¶ Predict the magnification factor as
sqrt(det(G))
for each point N in Xnew.
Parameters: - mean (bool) – whether to include the mean of the wishart embedding.
- covariance (bool) – whether to include the covariance of the wishart embedding.
- dimensions (array-like) – which dimensions of the input space to use [defaults to self.get_most_significant_input_dimensions()[:2]]
-
predict_noiseless(Xnew, full_cov=False, Y_metadata=None, kern=None)[source]¶ Convenience function to predict the underlying function of the GP (often referred to as f) without adding the likelihood variance on the prediction function.
This is most likely what you want to use for your predictions.
Parameters: - Xnew (np.ndarray (Nnew x self.input_dim)) – The points at which to make a prediction
- full_cov (bool) – whether to return the full covariance matrix, or just the diagonal
- Y_metadata – metadata about the predicting point to pass to the likelihood
- kern – The kernel to use for prediction (defaults to the model kern). this is useful for examining e.g. subprocesses.
Returns: - (mean, var):
mean: posterior mean, a Numpy array, Nnew x self.input_dim var: posterior variance, a Numpy array, Nnew x 1 if full_cov=False, Nnew x Nnew otherwise
If full_cov and self.input_dim > 1, the return shape of var is Nnew x Nnew x self.input_dim. If self.input_dim == 1, the return shape is Nnew x Nnew. This is to allow for different normalizations of the output dimensions.
Note: If you want the predictive quantiles (e.g. 95% confidence interval) use
predict_quantiles.
-
predict_quantiles(X, quantiles=(2.5, 97.5), Y_metadata=None, kern=None, likelihood=None)[source]¶ Get the predictive quantiles around the prediction at X
Parameters: - X (np.ndarray (Xnew x self.input_dim)) – The points at which to make a prediction
- quantiles (tuple) – tuple of quantiles, default is (2.5, 97.5) which is the 95% interval
- kern – optional kernel to use for prediction
Returns: list of quantiles for each X and predictive quantiles for interval combination
Return type: [np.ndarray (Xnew x self.output_dim), np.ndarray (Xnew x self.output_dim)]
-
predict_wishart_embedding(Xnew, kern=None, mean=True, covariance=True)[source]¶ Predict the wishart embedding G of the GP. This is the density of the input of the GP defined by the probabilistic function mapping f. G = J_mean.T*J_mean + output_dim*J_cov.
Parameters: Xnew (array-like) – The points at which to evaluate the magnification. :param
Kernkern: The kernel to use for the magnification.Supplying only a part of the learning kernel gives insights into the density of the specific kernel part of the input function. E.g. one can see how dense the linear part of a kernel is compared to the non-linear part etc.
-
predictive_gradients(Xnew, kern=None)[source]¶ Compute the derivatives of the predicted latent function with respect to X*
Given a set of points at which to predict X* (size [N*,Q]), compute the derivatives of the mean and variance. Resulting arrays are sized:
dmu_dX* – [N*, Q ,D], where D is the number of output in this GP (usually one).Note that this is not the same as computing the mean and variance of the derivative of the function!
dv_dX* – [N*, Q], (since all outputs have the same variance)Parameters: X (np.ndarray (Xnew x self.input_dim)) – The points at which to get the predictive gradients Returns: dmu_dX, dv_dX Return type: [np.ndarray (N*, Q ,D), np.ndarray (N*,Q) ]
-
set_XY(X=None, Y=None)[source]¶ Set the input / output data of the model This is useful if we wish to change our existing data but maintain the same model
Parameters: - X (np.ndarray) – input observations
- Y (np.ndarray) – output observations
-
to_dict(save_data=True)[source]¶ Convert the object into a json serializable dictionary. Note: It uses the private method _save_to_input_dict of the parent.
Parameters: save_data (boolean) – if true, it adds the training data self.X and self.Y to the dictionary Return dict: json serializable dictionary containing the needed information to instantiate the object
-
input_dim¶
-
num_data¶
GPy.core.gp_grid module¶
-
class
GpGrid(X, Y, kernel, likelihood, inference_method=None, name='gp grid', Y_metadata=None, normalizer=False)[source]¶ Bases:
GPy.core.gp.GPA GP model for Grid inputs
Parameters: - X (np.ndarray (num_data x input_dim)) – inputs
- likelihood (GPy.likelihood.(Gaussian | EP | Laplace)) – a likelihood instance, containing the observed data
- kernel (a GPy.kern.kern instance) – the kernel (covariance function). See link kernels
-
parameters_changed()[source]¶ Method that is called upon any changes to
Paramvariables within the model. In particular in the GP class this method reperforms inference, recalculating the posterior and log marginal likelihood and gradients of the modelWarning
This method is not designed to be called manually, the framework is set up to automatically call this method upon changes to parameters, if you call this method yourself, there may be unexpected consequences.
GPy.core.mapping module¶
-
class
Bijective_mapping(input_dim, output_dim, name='bijective_mapping')[source]¶ Bases:
GPy.core.mapping.MappingThis is a mapping that is bijective, i.e. you can go from X to f and also back from f to X. The inverse mapping is called g().
-
class
Mapping(input_dim, output_dim, name='mapping')[source]¶ Bases:
GPy.core.parameterization.parameterized.ParameterizedBase model for shared mapping behaviours
-
static
from_dict(input_dict)[source]¶ Instantiate an object of a derived class using the information in input_dict (built by the to_dict method of the derived class). More specifically, after reading the derived class from input_dict, it calls the method _build_from_input_dict of the derived class. Note: This method should not be overrided in the derived class. In case it is needed, please override _build_from_input_dict instate.
Parameters: input_dict (dict) – Dictionary with all the information needed to instantiate the object.
-
static
GPy.core.model module¶
-
class
Model(name)[source]¶ Bases:
paramz.model.Model,GPy.core.parameterization.priorizable.Priorizable-
static
from_dict(input_dict, data=None)[source]¶ Instantiate an object of a derived class using the information in input_dict (built by the to_dict method of the derived class). More specifically, after reading the derived class from input_dict, it calls the method _build_from_input_dict of the derived class. Note: This method should not be overrided in the derived class. In case it is needed, please override _build_from_input_dict instate.
Parameters: input_dict (dict) – Dictionary with all the information needed to instantiate the object.
-
objective_function()[source]¶ The objective function for the given algorithm.
This function is the true objective, which wants to be minimized. Note that all parameters are already set and in place, so you just need to return the objective function here.
For probabilistic models this is the negative log_likelihood (including the MAP prior), so we return it here. If your model is not probabilistic, just return your objective to minimize here!
-
objective_function_gradients()[source]¶ The gradients for the objective function for the given algorithm. The gradients are w.r.t. the negative objective function, as this framework works with negative log-likelihoods as a default.
You can find the gradient for the parameters in self.gradient at all times. This is the place, where gradients get stored for parameters.
This function is the true objective, which wants to be minimized. Note that all parameters are already set and in place, so you just need to return the gradient here.
For probabilistic models this is the gradient of the negative log_likelihood (including the MAP prior), so we return it here. If your model is not probabilistic, just return your negative gradient here!
-
randomize(rand_gen=None, *args, **kwargs)¶ Randomize the model. Make this draw from the prior if one exists, else draw from given random generator
Parameters: - rand_gen – np random number generator which takes args and kwargs
- loc (flaot) – loc parameter for random number generator
- scale (float) – scale parameter for random number generator
- kwargs (args,) – will be passed through to random number generator
-
static
GPy.core.sparse_gp module¶
-
class
SparseGP(X, Y, Z, kernel, likelihood, mean_function=None, X_variance=None, inference_method=None, name='sparse gp', Y_metadata=None, normalizer=False)[source]¶ Bases:
GPy.core.gp.GPA general purpose Sparse GP model
This model allows (approximate) inference using variational DTC or FITC (Gaussian likelihoods) as well as non-conjugate sparse methods based on these.
This is not for missing data, as the implementation for missing data involves some inefficient optimization routine decisions. See missing data SparseGP implementation in py:class:’~GPy.models.sparse_gp_minibatch.SparseGPMiniBatch’.
Parameters: - X (np.ndarray (num_data x input_dim)) – inputs
- likelihood (GPy.likelihood.(Gaussian | EP | Laplace)) – a likelihood instance, containing the observed data
- kernel (a GPy.kern.kern instance) – the kernel (covariance function). See link kernels
- X_variance (np.ndarray (num_data x input_dim) | None) – The uncertainty in the measurements of X (Gaussian variance)
- Z (np.ndarray (num_inducing x input_dim)) – inducing inputs
- num_inducing (int) – Number of inducing points (optional, default 10. Ignored if Z is not None)
-
parameters_changed()[source]¶ Method that is called upon any changes to
Paramvariables within the model. In particular in the GP class this method re-performs inference, recalculating the posterior and log marginal likelihood and gradients of the modelWarning
This method is not designed to be called manually, the framework is set up to automatically call this method upon changes to parameters, if you call this method yourself, there may be unexpected consequences.
-
plot_inducing(visible_dims=None, projection='2d', label='inducing', legend=True, **plot_kwargs)¶ Plot the inducing inputs of a sparse gp model
Parameters: - visible_dims (array-like) – an array specifying the input dimensions to plot (maximum two)
- plot_kwargs (kwargs) – keyword arguments for the plotting library
GPy.core.sparse_gp_mpi module¶
-
class
SparseGP_MPI(X, Y, Z, kernel, likelihood, variational_prior=None, mean_function=None, inference_method=None, name='sparse gp', Y_metadata=None, mpi_comm=None, normalizer=False)[source]¶ Bases:
GPy.core.sparse_gp.SparseGPA general purpose Sparse GP model with MPI parallelization support
This model allows (approximate) inference using variational DTC or FITC (Gaussian likelihoods) as well as non-conjugate sparse methods based on these.
Parameters: - X (np.ndarray (num_data x input_dim)) – inputs
- likelihood (GPy.likelihood.(Gaussian | EP | Laplace)) – a likelihood instance, containing the observed data
- kernel (a GPy.kern.kern instance) – the kernel (covariance function). See link kernels
- X_variance (np.ndarray (num_data x input_dim) | None) – The uncertainty in the measurements of X (Gaussian variance)
- Z (np.ndarray (num_inducing x input_dim)) – inducing inputs
- num_inducing (int) – Number of inducing points (optional, default 10. Ignored if Z is not None)
- mpi_comm (mpi4py.MPI.Intracomm) – The communication group of MPI, e.g. mpi4py.MPI.COMM_WORLD
-
optimize(optimizer=None, start=None, **kwargs)[source]¶ Optimize the model using self.log_likelihood and self.log_likelihood_gradient, as well as self.priors. kwargs are passed to the optimizer. They can be:
Parameters: - max_iters (int) – maximum number of function evaluations
- messages (bool) – whether to display during optimisation
- optimizer (string) – which optimizer to use (defaults to self.preferred optimizer), a range of optimisers can be found in :module:`~GPy.inference.optimization`, they include ‘scg’, ‘lbfgs’, ‘tnc’.
- ipython_notebook (bool) – whether to use ipython notebook widgets or not.
- clear_after_finish (bool) – if in ipython notebook, we can clear the widgets after optimization.
-
parameters_changed()[source]¶ Method that is called upon any changes to
Paramvariables within the model. In particular in the GP class this method re-performs inference, recalculating the posterior and log marginal likelihood and gradients of the modelWarning
This method is not designed to be called manually, the framework is set up to automatically call this method upon changes to parameters, if you call this method yourself, there may be unexpected consequences.
-
optimizer_array¶ Array for the optimizer to work on. This array always lives in the space for the optimizer. Thus, it is untransformed, going from Transformations.
Setting this array, will make sure the transformed parameters for this model will be set accordingly. It has to be set with an array, retrieved from this method, as e.g. fixing will resize the array.
The optimizer should only interfere with this array, such that transformations are secured.
GPy.core.svgp module¶
-
class
SVGP(X, Y, Z, kernel, likelihood, mean_function=None, name='SVGP', Y_metadata=None, batchsize=None, num_latent_functions=None)[source]¶ Bases:
GPy.core.sparse_gp.SparseGPStochastic Variational GP.
For Gaussian Likelihoods, this implements
Gaussian Processes for Big data, Hensman, Fusi and Lawrence, UAI 2013,
But without natural gradients. We’ll use the lower-triangluar representation of the covariance matrix to ensure positive-definiteness.
For Non Gaussian Likelihoods, this implements
Hensman, Matthews and Ghahramani, Scalable Variational GP Classification, ArXiv 1411.2005
-
new_batch()[source]¶ Return a new batch of X and Y by taking a chunk of data from the complete X and Y
-
parameters_changed()[source]¶ Method that is called upon any changes to
Paramvariables within the model. In particular in the GP class this method re-performs inference, recalculating the posterior and log marginal likelihood and gradients of the modelWarning
This method is not designed to be called manually, the framework is set up to automatically call this method upon changes to parameters, if you call this method yourself, there may be unexpected consequences.
-